Feb 6, 2023 10:25:20 AM
How to purify your peptide using mass directed flash chromatography
By Elizabeth Denton

We've all used mass spectrometry to characterize our synthetic peptides. It's often used to confirm that the peptide was in fact synthesized, then again as part of the purification process to make sure that we're collecting the correct peak. But how many of you had the opportunity to use in-line mass spectrometry as an integral component during the purification itself?
In today's post, I'll highlight some of the advantages to using in-line mass mass spectrometry for purification of peptides using reversed phase flash chromatography.
Flash chromatography has become more commonplace in the peptide lab, complementing existing HPLC purification capability or even in some cases replacing it altogether. The biggest challenge to overcome when switching from an HPLC-mediated purification strategy to a flash-mediated purification strategy is the decreased resolution due to the larger stationary phase. I have talked about several different strategies that can help overcome this loss of resolution in the past (changing mobile phase additives, using a step gradient, etc). But one area I haven't touched on much is the use of in-line mass directed purification.
The most significant advantages to using an in-line mass detector for your purification is efficiency and confidence. The real-time feedback reduces the amount of time spent analyzing each and every collected fraction to determine where in fact the product of interest may lie within the chromatogram.
Nowadays, I'm using my Biotage® Isolera Dalton 2000 mass directed flash purification system to purify my peptides with a couple different approaches. It's important to note that the Isolera Dalton 2000 can be mass triggered using both the total ion chromatogram (TIC) for positive and negative ions simultaneously if you are unsure of the most abundant masses in the sample, or the user can specifically define four unique extracted ions (XIC) for analysis, or both methods at once!
In a preliminary "scouting" type injection, I like to monitor my chromatography with the mass information, Figure 1. In this scenario, the mass information selected is still displayed on the chromatogram, but the actual fraction collection is triggered by the UV detector. This allows for the greatest sensitivity and sample recovery as the threshold for the UV fractionation to can be adjusted by the user. 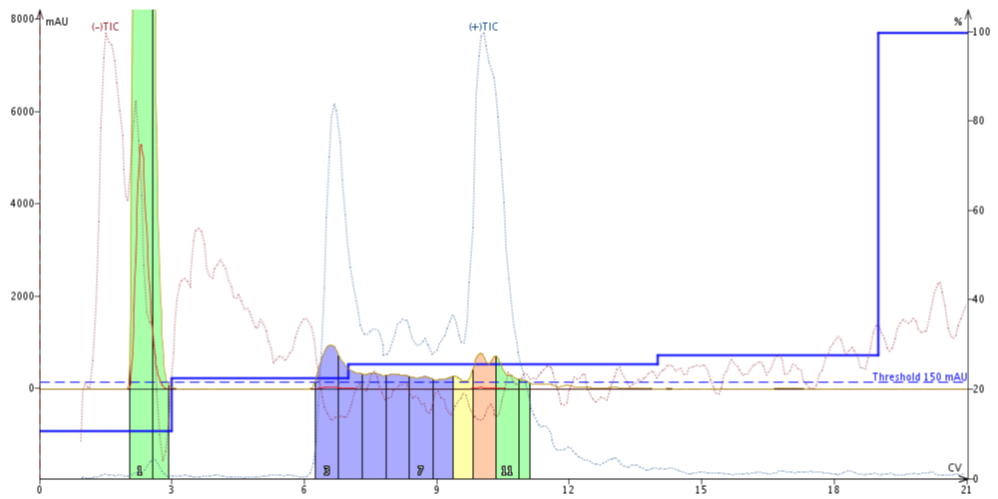
Figure 1. Mass guided flash chromatography using only the positive and negative total ion chromatogram. The result can be further interrogated to identify the specific mass ions that give rise to the TIC peaks visible.
This strategy also allows me to select the most abundant masses detected by the Dalton 2000 mass spectrometer. I have observed that the precise molecular weight can vary slightly between individual mass spectrometers, Figure 2. These small differences can absolutely impact the sample recovery for future purifications so it is critical to select the most abundant ions detected by this spectrometer.
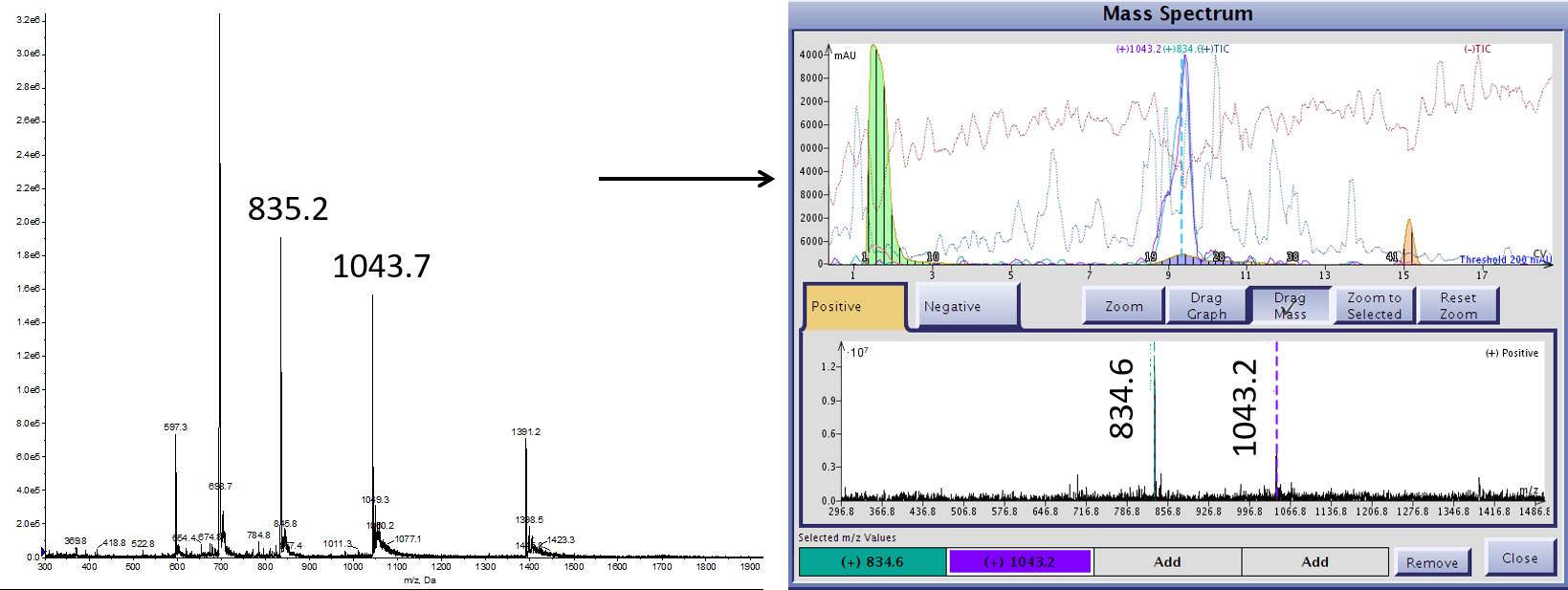
Figure 2. Follow up mass analysis indicates subtle differences in the predominant mass species observed using the Isolera Dalton 2000 (right) and a more standard analytical MS (left) for the same crude peptide sample.
For larger quantities of peptide to be purified, I often trigger fraction collection using the mass spectrometer as one would expect. I have used this strategy to triple my typical loading level (300 mg crude loaded onto a 10 g C18 cartridge!!!) while still achieving high purity and high sample recovery for a relatively clean peptide, Figure 3.
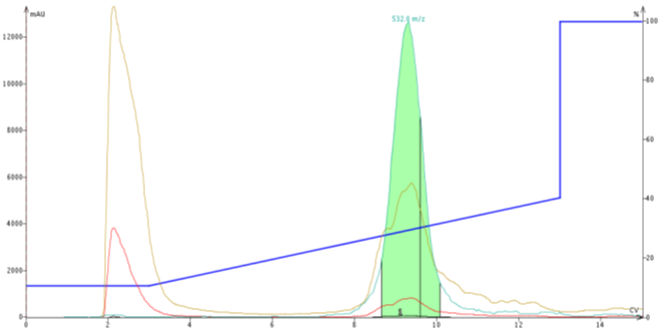
Figure 3. Purification of 300 mg crude ACP peptide using mass-directed fractionation. The alternative sensitivity increased final sample purity from 31% to >92% and a full 3% sample load relative to cartridge packing content.
More often than not though, I use the four XIC masses to assist in the purification. I will typically assign two of the four masses to ions corresponding to the peptide of interest and use the other two available XIC masses for closely eluting impurities typically not resovable by the UV detector alone and increasing the final peptide purity, Figure 4.
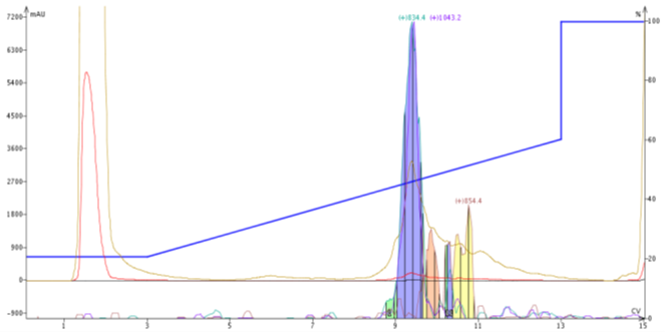
Figure 4. Separation of a 20 amino acid peptide from a 19 amino acid impurity is possible using mass-directed and specified extracted ions, but are not resolvable using only UV (tan line).
This approach is effective primarily because of the way the Isolera Dalton 2000 software interprets the XIC data. The software was originally designed for small molecules, so it thinks that each XIC mass corresponds to a "different" compound. For example, as one XIC mass begins to decrease in intensity (near the end of compound elution), a new fraction will be triggered when a second XIC mass begins to increase in intensity (beginning of a new peak), Figure 5.
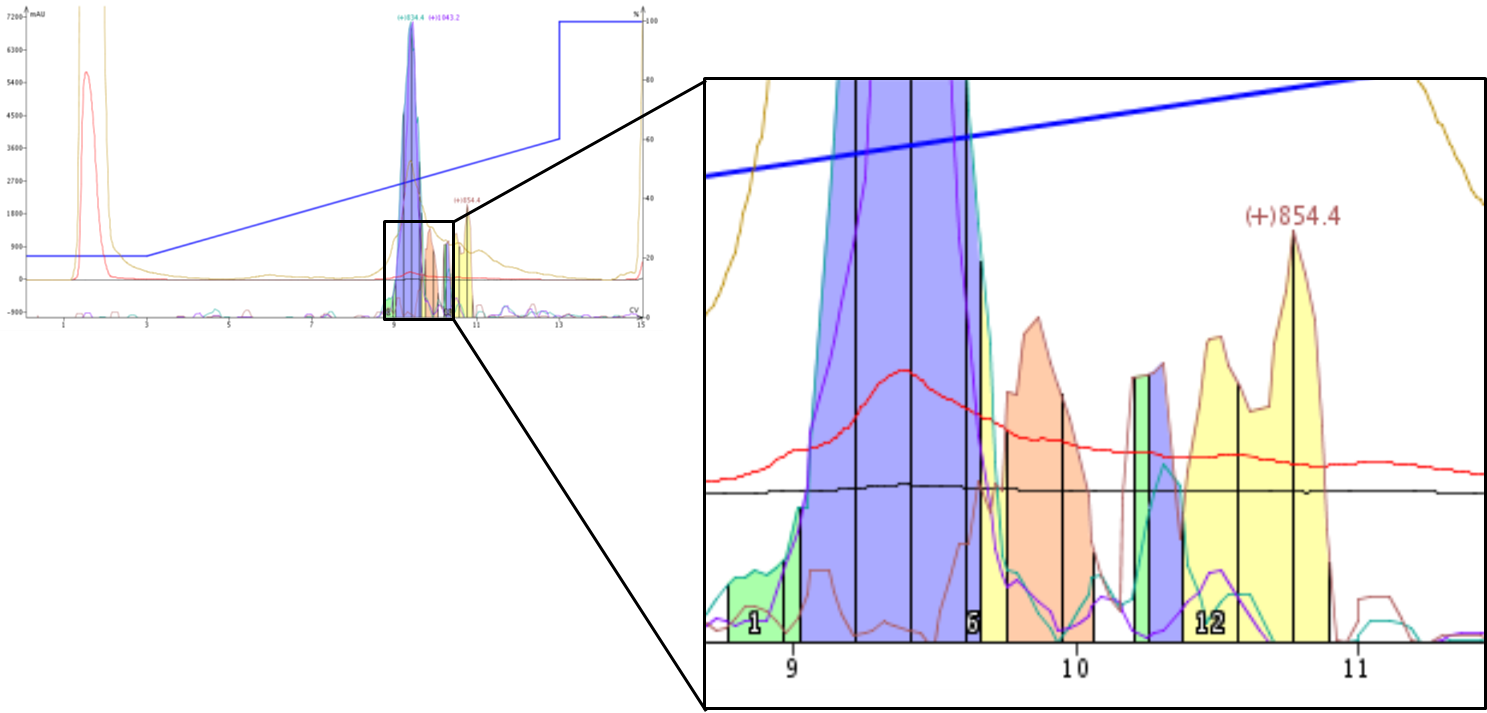
Figure 5. Zoomed view demonstrates separation of single amino acid deletions with carefully defined extract ion masses. A new fraction is collected (yellow center) when the desired peptide mass intensities (purple and teal lines) is sufficiently decreased and the new impurity mass intensity begins to increase (red line).
Using mass-directed, or even simply mass-guided flash purification has not only reduced my purification time, it has also increased my confidence that the fractions I've isolated and move forward for evaporation contain my desired peptide in high purity.
If you would like to learn more about the advantages of flash chromatography for peptide purification and the strategies that enable recovery of highly pure peptides, click the link below.

