Sep 12, 2023 12:55:46 AM
Can I make head-to-tail macrocycles on-resin?
By Austin Schlirf

Macrocyclization of peptides has arisen as a new strategy to create proteolytically stable, membrane permeable drug candidates while maintaining target specificity. To perform this cyclization, lactam and disulfide bridges are simple orthogonal techniques that allow for these macrocyclic structures to be synthesized on-resin. Nature has inspired a third type of cyclic peptide in the form of head-to-tail peptide macrocycles. These head-to-tail cyclic peptide are becoming a unique structure that for typical solid phase peptide synthesis (SPPS) is impossible due to the resin bound C-terminus. I asked, what would happen if I could free up the C-terminus while keeping my peptides on-resin? Here I present the strategy that worked for me to free up the C-terminus and methods for cyclizing your peptides.
I began by thinking of ways to link my peptide to the resin in a way that allows the C-terminus to remain free. The obvious choice here was to give myself two COOH groups, so using Asp and Glu allows for that to happen. I regularly use Rink-amide resins and working with and cleaving from these resins is typically fairly straightforward. Yes, that gives you an amide on your COOH bound sidechain, but the end product is still a canonical amino acid (Gln/Asn) and I can work with that for simplicity's sake.
So, I started off with the strategy of linking the side chain of Glu residues to a Rink amide polystyrene resin knowing I would end up with a final product containing Gln. So, what should my sequence be? Commonly, macrocyclic peptides are ~6-15 residues in length, but library screens can generate lots of different sequences. So, lets choose something that, from a cyclization perspective, will make things a bit difficult. Below in Figure 1 is the sequence I chose. It has some fairly bulky groups, charged moieties, and aliphatic side chains that all might be found in peptides of interest.
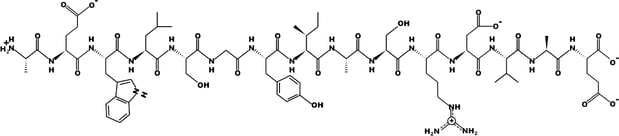
H-AEWLSGYIASRDVAD-OH
Figure 1. Linear peptide sequence
I used traditional FMOC protecting groups except for my Asp and Glu residues. Here I protected the main chain carboxylic acid with an allyl protecting group which is orthogonally removed and typically works well for me. I synthesized my linear peptides using our standard methods on a Biotage® Initiator+ Alstra™ peptide synthesizer. Here my linear Glu-linked peptide was not as clean as I would prefer but, in the end, I would be comparing linear peptide to cyclized product so I can work with this, Figure 2.
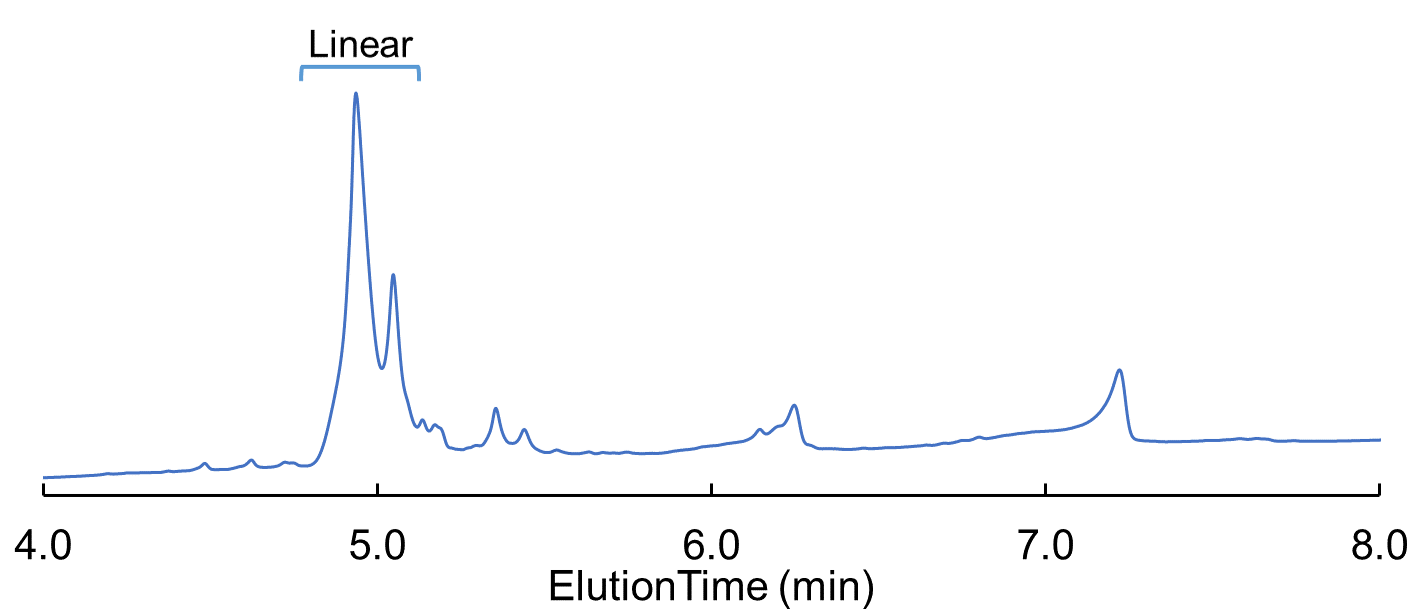
Figure 2: Linear Glu-linked peptide HPLC chromatogram
After removing the allyl protecting group, I tried my first cyclization and decided to really hit it over the head with a hammer to make sure it worked. I did a double DIC/Oxyma coupling and had longer reaction times (Coupling 1 - 50°C, 30min; Coupling 2 - 50°C, 60min); to my surprise I was successful, Figure 3.
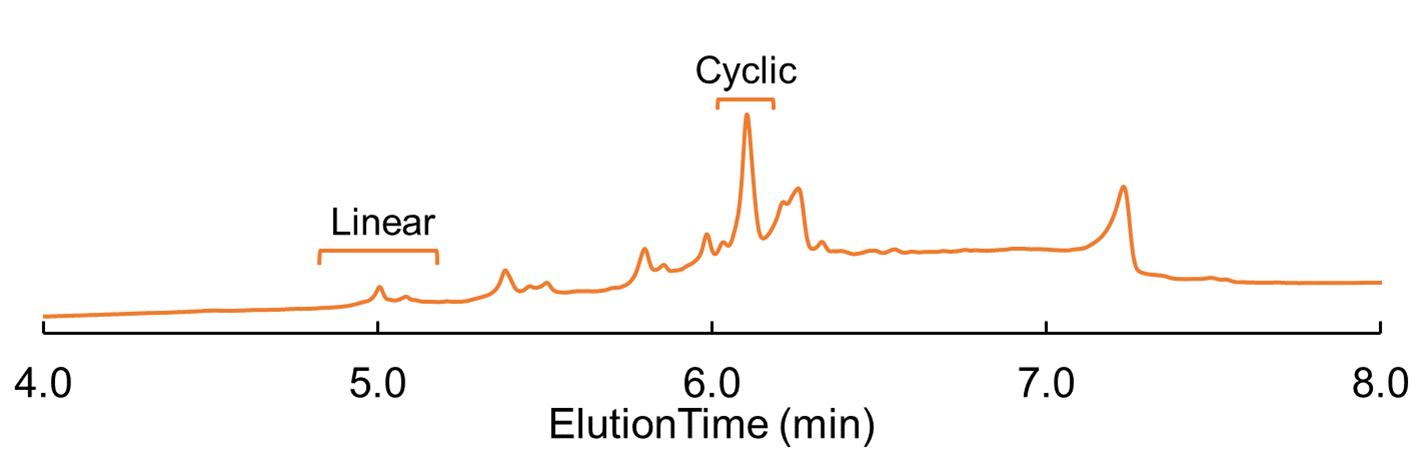
Figure 3: Cyclic Glu-linked peptide HPLC chromatogram; cyclized using double coupling at 50°C for 30 min. and 60 min.
However, not everyone likes long coupling times and double couplings, so I wanted to try a shorter single coupling (10 min) as well with higher temperature (75°C), Figure 4. In retrospect, there is nothing wrong with a longer reaction time here as this is going for your final product and you want as much of it as you can especially if you are doing this in a larger batch. So why not go low and slow?
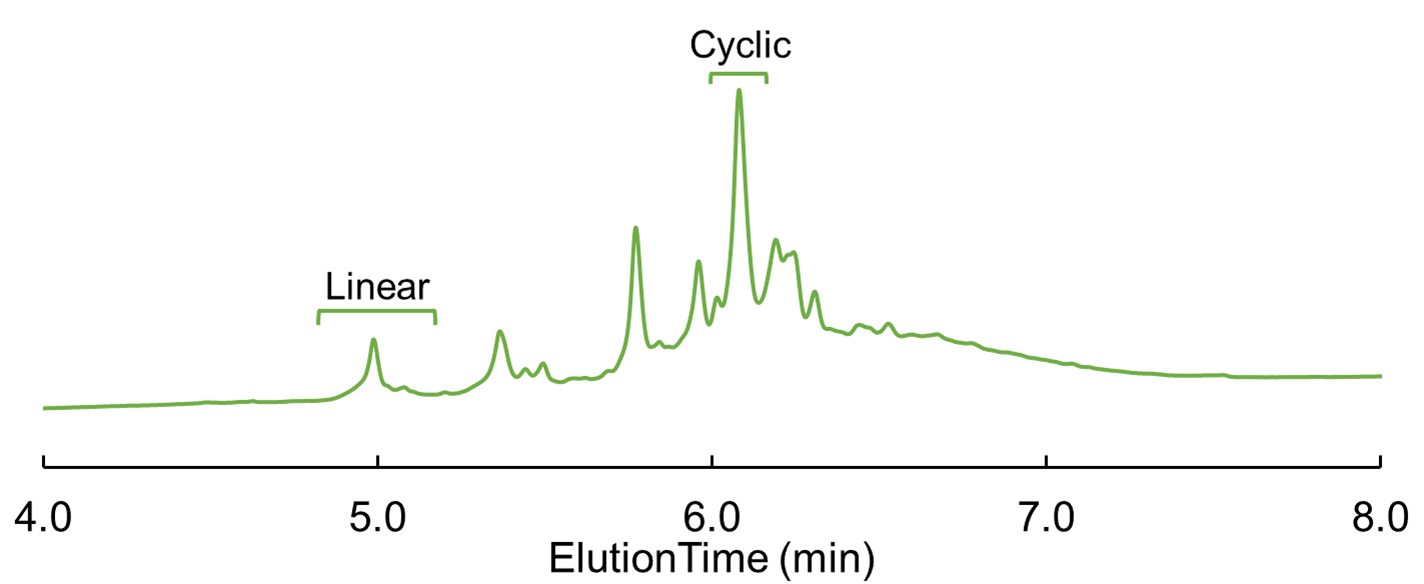
Figure 4: Cyclic Glu-linked peptide HPLC chromatogram; cyclized using single coupling at 75°C for 10 min.
Not too shabby right? Overall, it seems that we can make these cyclic products with this strategy, and it is fairly straightforward.
| Resin Bound AA | Coupling Scheme | Temperature (°C) | Coupling 1 (min) | Coupling 2 (min) | % Cyclic in Crude | % Linear in crude |
| Glu | Double | 50 | 30 | 60 | 28.0 | 4.0 |
| Glu | Double | 50 | 30 | 10 | 22.5 | 4.5 |
| Glu | Single | 75 | 10 | - | 22.8 | 4.3 |
Cyclic peptides come in all shapes and sizes, and this is the beginning of what may be a possible technique for simplifying those head-to-tail cycles for me. I definitely want to try a range of peptide lengths (as shorter may actually be harder), resin loading (0.3 mmol/g vs 0.49 mmol/g) to reduce the potential of dimerization, using wang instead of rink-amide linker or switch up my coupling reagents as not everyone uses DIC/Oxyma.
What strategies have you tried to make your cyclic peptides? Is there something you'd like to see attempted by yours truly? If you want to read more on cyclic peptide, see the link below!


